Help
This page contains help and guidelines on how to use the NABh Classifier effectively.
Content
- 1. What is NABhClassifier?
- 2. What can NABhClassifier do?
- 3. How to Use NABhClassifier
- 4. Parameters and Outputs
- 5. Result Scenarios
- 6. How to cite us
1. What is NABhClassifier?
NABhClassifier is a web-based tool designed to identify helical sequences in proteins with potential nucleic acid-binding capabilities (DNA and RNA). Developed using machine learning models, NABhClassifier provides a fast and accurate solution for detecting small helical domains capable of interacting with nucleic acids, facilitating research in biotechnology and medicine.
2. What can NABhClassifier do?
NABhClassifier can:
- Identify alpha helices in protein sequences.
- Classify helices with potential nucleic acid-binding capabilities.
- Generate a NABh Index, which combines the results of eight machine learning models to provide a confidence score.
- Analyze protein sequences in seconds with an intuitive and easy-to-use interface.
3. How to Use NABhClassifier
Follow these steps to use NABhClassifier:
-
Input the Sequence: In the input field, paste the protein sequence in FASTA format.
>Q8CY87 MVVFTGSTVEEAIQKGLKELDIPRMKAHIKVISREKKGFLGLFGKKPAQVDIEAISETTV VKANQQVVKGVPKKINDLNEPVKTVSEETVDLGHVVDAIKKIEEEGQGISDEVKAEILKH ERHASTILEETGHIEILNELQIEEAMREEAGADDLETEQDQAESQELEDLGLKVETNFDI EQVATEVMAYVQTIIDDMDVEATLSNDYNRRSINLQIDTNEPGRIIGYHGKVLKALQLLA QNYLYNRYSRTFYVTINVNDYVEHRAEVLQTYAQKLATRVLEEGRSHKTDPMSNSERKII HRIISRMDGVTSYSEGDEPNRYVVVDTE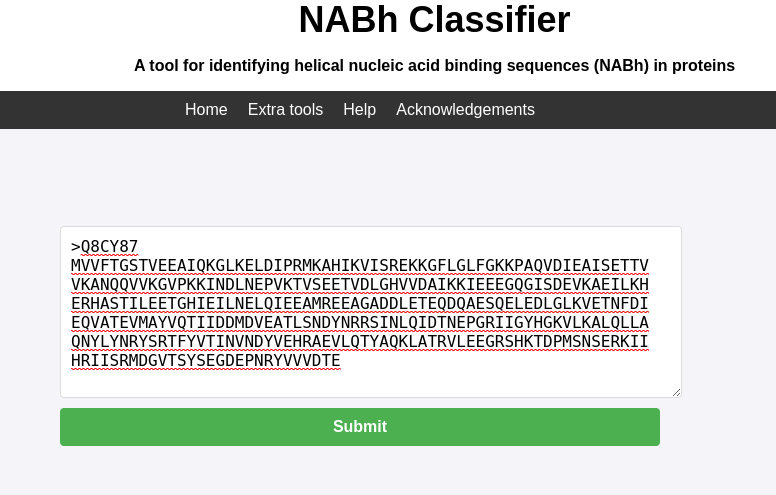
- Submit the Analysis: Click "Submit" to start the analysis. The server will process the sequence automatically.
-
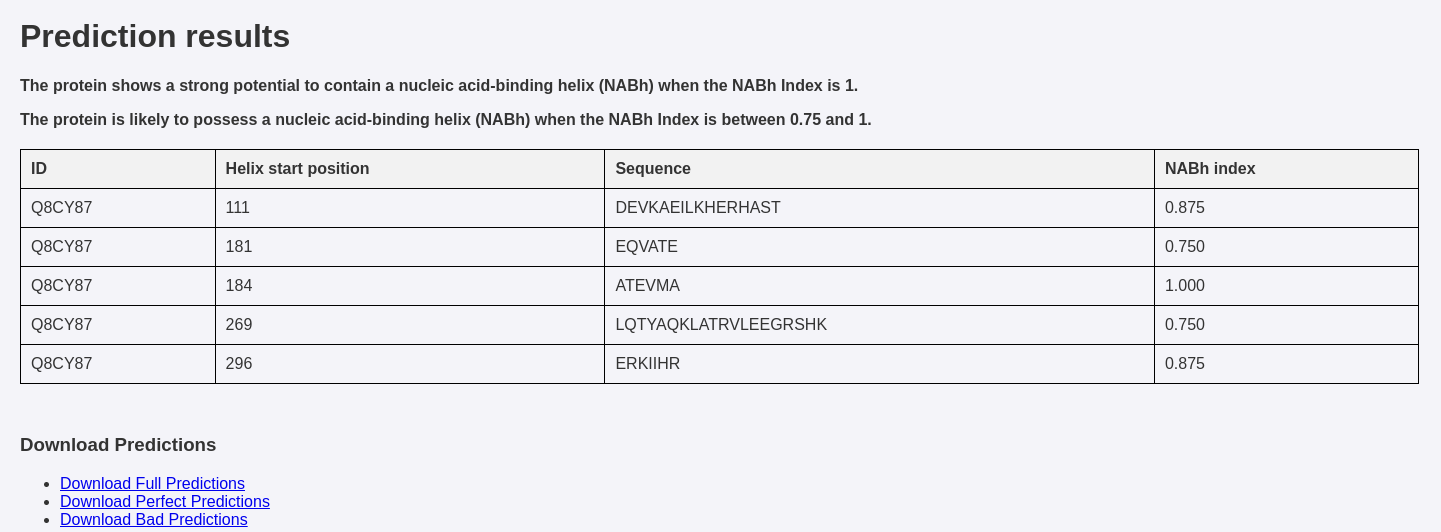
View and Download Results: After analysis, the results will be displayed on the results page, showing the identified helices and their NABh Index scores. You can also download the results in CSV format for further analysis.
4. Parameters and Outputs
NABhClassifier generates the following results:
- NABh Index: A consensus score ranging from 0 to 1, where:
- 1.0: Full agreement among models (high confidence).
- 0.75-0.875: Partial agreement (moderate probability).
- Below 0.75: Low confidence (not recommended for practical use).
- Helix Position: The location of helices in the protein sequence.
- Helix Sequence: The amino acid sequence of the identified helix.
5. Result Scenarios
Here are some common scenarios you may encounter when using NABhClassifier and how to interpret them:
-
Scenario 1: The input sequence is not recognized.
- Explanation: This usually happens when the sequence is not in FASTA format or contains non-standard amino acids.
- Solution: Ensure the sequence is in FASTA format and contains only standard amino acids.
-
Scenario 2: No helices were identified.
- Explanation: This occurs when the algorithm does not find enough residues to form a helix.
-
Scenario 3: Results with low NABh Index.
- Explanation: This indicates that the identified helices have a low probability of binding to nucleic acids.
-
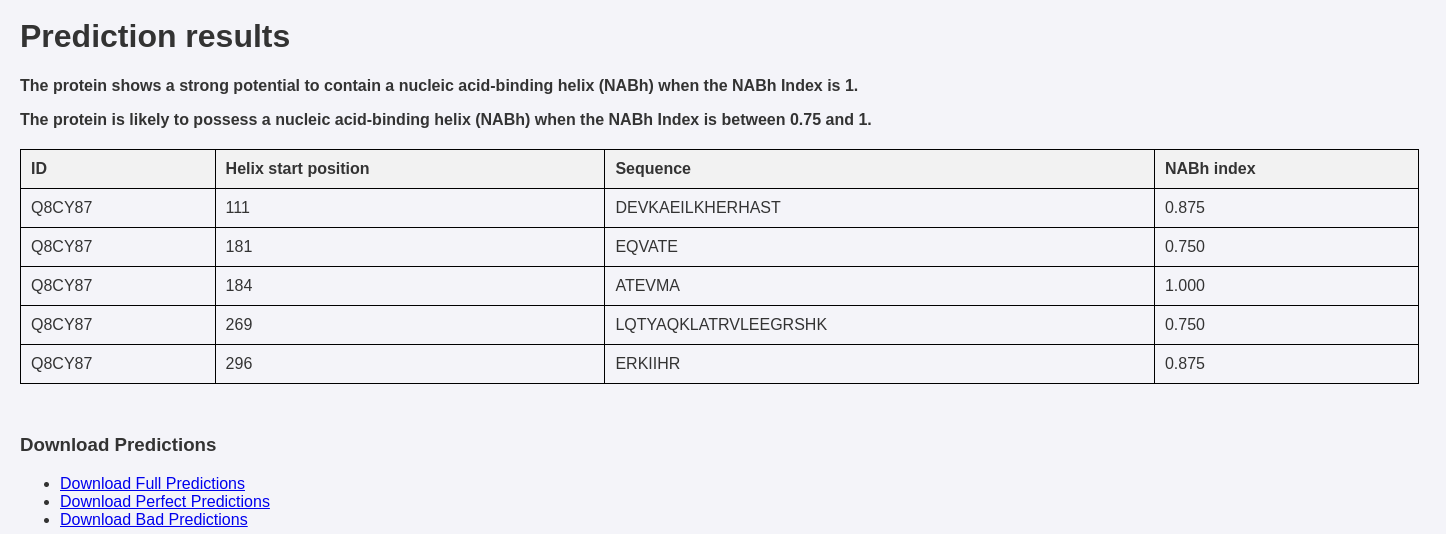
Scenario 4: Results with high NABh Index.
- Explanation: This indicates that the identified helices have a high probability of binding to nucleic acids. These results are reliable and can be used for further analysis or experimental validation.
6. How to cite us
If you use NABhClassifier in your research, please cite the following article:
NABhClassifier Server: A Tool for the Identification of Helical Nucleic Acid-Binding Sequences in Proteins.
Rogerio Margis, Iara Macedo, Nureyev F. Rodrigues, Mateus Dias-Oliveira, Fernanda Lazzarotto, Diego Trindade de Souza, and Geancarlo Zanatta.
Journal of Chemical Information and Modeling 2025, 65 (5), 2361-2367.
DOI: 10.1021/acs.jcim.4c02244.